Prions
"Amyloidogenic domains, prions and structural inheritance: rudiments of early life or recent acquisition?", by Y. O. Chernoff, Current Opinion in Chemical Biology, 2004, 8, 665 - 671
Fibre-like protein polymerization is involved in the formation of biological structures. It is possible that such structural inheritance played a role in the early steps of biological compartmentalization, eventually leading to the formation of primordial cells. In contrast to in vitro amyloid a, b formation, in vivo propagation of yeast prions requires cellular co-factors such as the yeast chaperone Hsp 104 (heat shock protein). Hsp 104 was hypothesized to work in prion propagation by shearing large polymers into oligomeric seeds, initiating new rounds of amyloid polymerization (see the Figure below). This may lend itself to a grammar, as well.
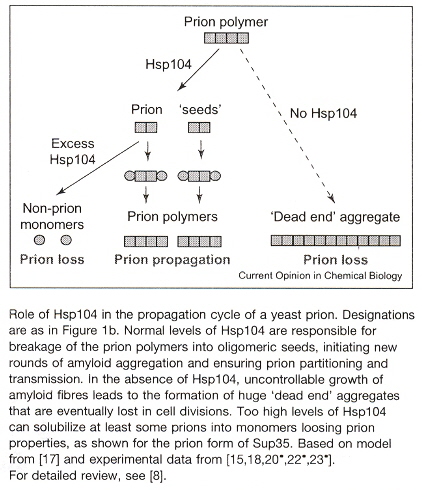
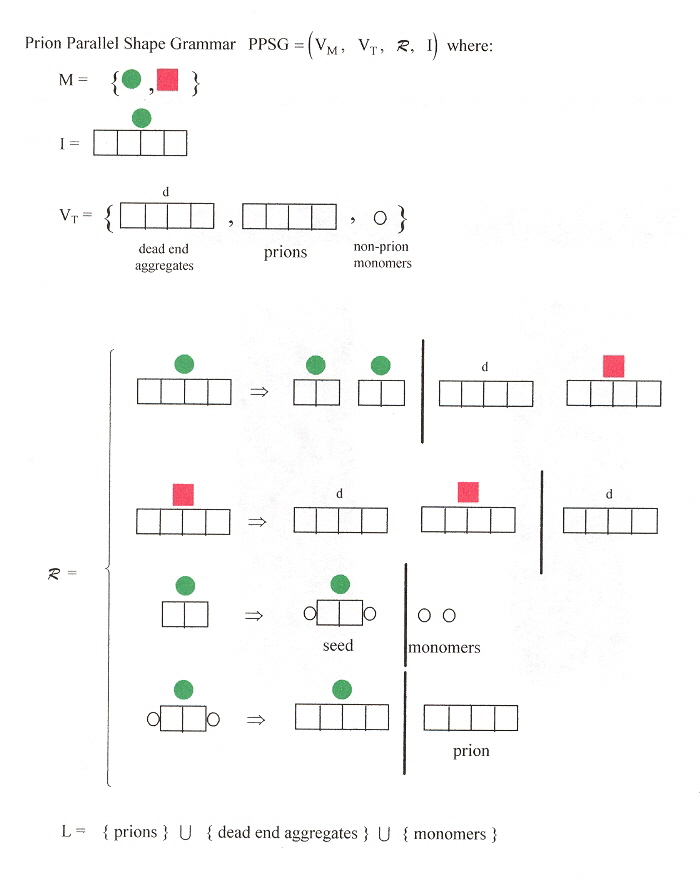
a "Template-Directed Self-Assembly and Growth of Ijsulin Amyloid Fibrils", by C. Ha, C. B. Pak, Biotechnology and Bioengineering, June 30 2005, 90, 7, 848 - 855
Amyloid is a term first introduced by Rudolph Virchow in 1854, and refers to protein aggregates with a fibrillar morphology, cross β-sheet secondary structure, bifringent staining with Congo red dye, and is insoluble in aqueous solution. Amyloid formation takes place dominatly in the proximity of membranes, and is notably accelerated in the presence of preformed plaques (amyloid templates). For just this reason, Esler et al developed an "artificial" template.
b "Alzheimer's disease amyloid propagation by a template-dependent dock–lock mechanism", by W. P. Esler, E. R. Stimson, J. M. Jennings, H. V. Vinters, J. R. Ghilardi, J. P. Lee, P. W. Mantyh, J. E. Maggio, Biochemistry, 2000, 39, 6288 - 6295
"Strain-specified characteristics of mouse synthetic prions", by G. Legname, H-O. B. Nguyen, I. V. Baskakov, F. E. Cohen, S. J. De Armond, S. B. Prusiner, Proceedings of the National Academy of Sciences U. S., Feb. 8 2005, 102, 6, 2168 - 2173
A synthetic mouse prion strain MoSP1 was created as N-terminally truncated MoPrP(Δ23-88). The criteria used to determine that this synthetic prion is new include the following.
- Neuropathological changes that are specific, such as large vacuoles in the cerebellar gray matter and brainstem. Other prions have widespread vacuoles including areas such as the neocortex, hippocampus, thalmus, brainstem, and cerebellar white matter.
- Conformation stability of MoSP1Sc to increasing concentrations of guanidine HCl (GdnHCL)
- Incubation time profiles until the effects of MoSP1 appear (516±27 days, on first passage; 258±25 days, on second passage in mice)
"Biology of infectious proteins: lessons from yeast prions", by M. Burwinkel, N. Holtkamp, M. Baier, Lancet, Feb. 12 2005, 365, 9459, 572
Sup35p is a yeast prion that normally terminates translation at stop codons. It can also adopt a prion conformation. However, Sup35p protein does not function in prion-infected yeast, as the ribosomal machinery reads beyond stop codons. By translating genetic information normally hidden behind stop codons, infected yeast cells are equipped with new phenotypes that are occasionally advantageous. Proteins with altered properties thus contribute to the diversity of yeast populations. This enhanced phenotypic diversity facilitates the yeast's adaptation to environmental changes. When exposed to toxic environments that would normally prevent growth of uninfected yeast, prion-infected yeast survive: yeast thus benefits from increased biological diversity. However, prions are non-chromosomally inherited. If the prion infection favours a cell in particular circumstances, the cell passes on this trait to its progeny. Prion infection can be lost spontaneously, but the advantages may still be maintained genetically by other mechanisms such as genetic reassortment or new mutations (benefits can be prion independent). Thus Sup35p prions promote evolutionary adaptation.
"A scientific revolution?", by A. E. Bussard, European Molecular Biology Organization, Aug. 2005, 6, 8, 691 - 694
The point of view expressed in this paper is that prions may challenge the central dogma of molecular biology. Specifically, may there be other mechanisms of heredity in living organisms than that which is encoded through DNA? There is growing evidence that hereditary information may be transferred genetically through the prion. The author points out that at the 1967 Cold Spring Harbor Symposium on Quantitative Biology, a discussion took place with F. Crick about the problem of the irreversibility of information transfer from nucleic acids to proteins. Although F. Crick accepted reversibility between DNA and RNA through retroviruses, the irreversibility of information transfer from RNA to proteins was considered absolute: "Nature could not proceed in another way." Thus the view was accepted that proteins encode the information stored in DNA. However, the idea that proteins could transmit information did not disappear (C. Gajdusek), this idea finally being accepted with Prusiner's theory about Prions. Pruisner's theory of Prions was a revolution against the dogma that only viruses and bacteria (that carry nucleic acids) could be infectious. The work by Lindquist on yeast prions showed that prion domains in some proteins act as molecular switches that activate or deactivate some proteins. Lindquist showed that prions are non-Mendelian genetic elements that produce new phenotypes, better adapted to the environment. Obviously, the tertiary structure and function of a protein is not determined solely by its amino-acid sequence. As inherited variance is apparently not restricted to DNA but could also be caused by protein-based genetic elements, some now wonder about reviving Lamarck's idea that the environment triggers adaptive structural and physiological changes in the organism. The prion model also puts in doubt the notion that cloned animals are genetically identical to their genome donors.
It is unfortunate that researchers dealing with molecular genetics (including prions) do not appear to be interested in or knowledgeable enough to deal with Mathematical Linguistics. An analysis of prions from the viewpoint of Mathematical Linguistics could prove to be of productive value. It is the viewpoint of the author of "Emergent Computation: Emphasizing Bioinformatics" that only the reported views of scientists working with molecular genetics be presented, thus no attempt will be made to analyze prions from the viewpoint of Mathematical Linguistics.
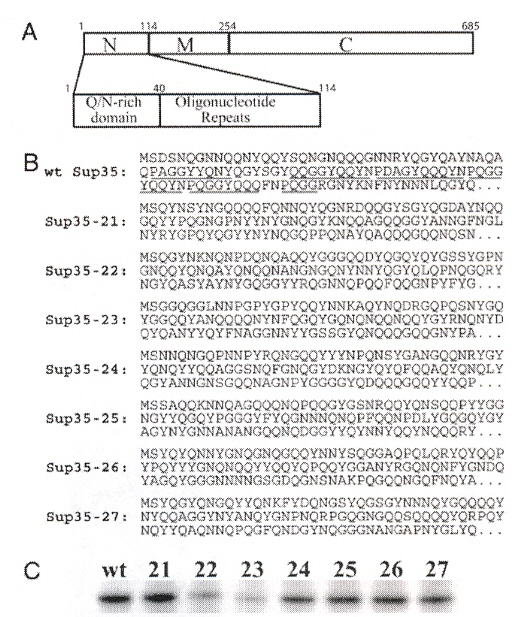
"Primary sequence independence for prion formation", by E. D. Ross, H. K. Edskes, M. J. Terry, R. B. Wickner, Proceedings of the National Academy of Sciences of the U.S., Sept. 6 2005, 102, 36, 12825 - 12830
Ten proteins are modified to investigate their ability to cause prion infection. These proteins include Ure2p, Ure2-21p, wt Sup35, Sup35-21 through Sup35-27. Ure2p and Sup35p are composed of a Prion Domain (PD) that terminates in amino acid N (asparginine) and a functional domain (7-11) that terminates in C (cysteine). Ure2p has a PD composed of residues 1-89 or 1-65. Ure2p is then followed by the functional domain residues 90-354. Sup35p has a PD composed of residues 1-114, followed by domain M (residues 115-253), and the remaining functional domain residues at 254-685. Fig. 1 describes Ure2p, Fig. 2 describes Ure2-21p, and Fig. 3 describes Sup35. In this paper, the amino acid residues of these proteins are randomly ordered to determine the effect on prion infection (protein misfolding). The results indicate that the PD infectious activity does not depend upon residue order, rather it is the residue composition.
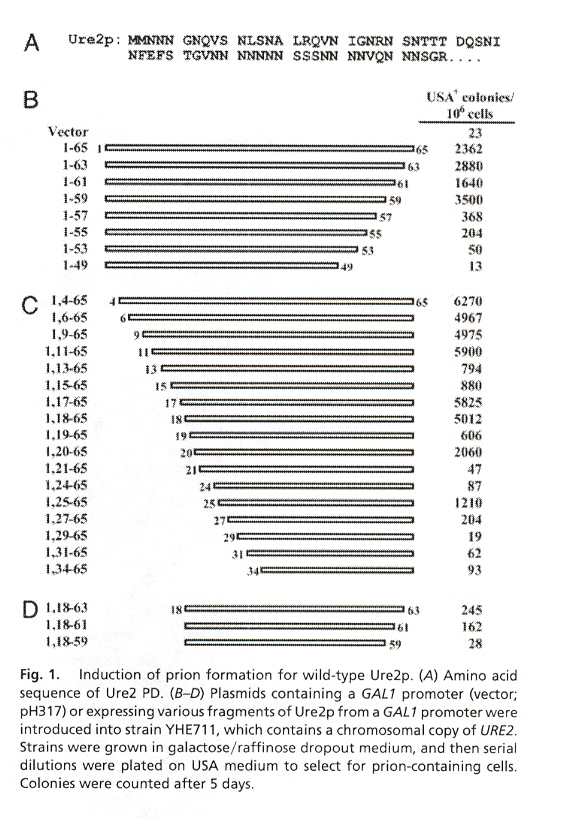
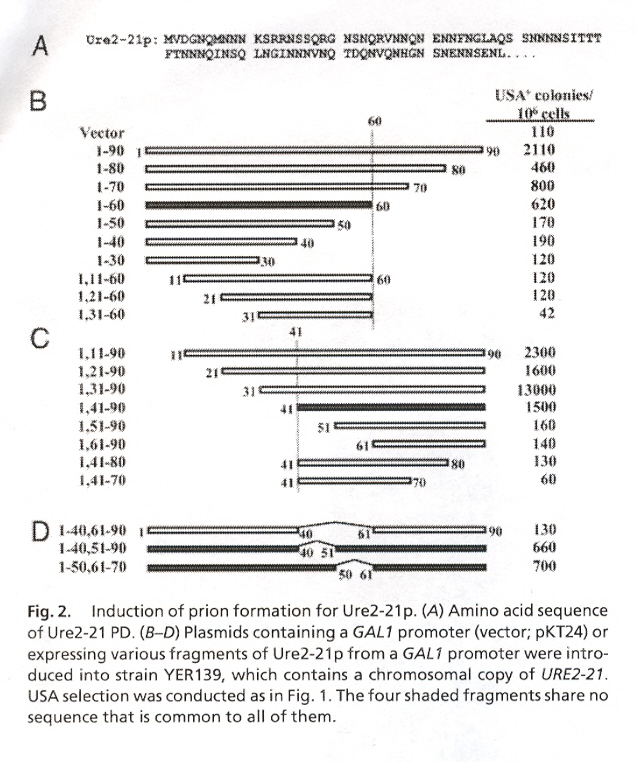
"Copper-metalated peptide palindrome derived from prion octarepeat: synthesis, aggregation, and oxidative transformations", by C. Madhavaiah, S. Verma, Bioorganic & Medicinal Chemistry, May 2 2005, 13, 9, 3241 - 4248
Normal prion PrPC binds copper at its N-terminal domain. Four octarepeats in the N-terminal of PHGGGWGO selectively binds Cu++ over other divalent metal ions in 1:1 stoichiometry. Such a copper-bound octapeptide region of prion protein exhibits superoxide dismutase activity, hence serving as a possible defense against oxidative stress in the brain. Other researchers a have demonstrated the ability of copper coordination to induce β–sheet in prion protein thus raising the possibility of Cu++ acting as a trigger for prion misfolding.
a "Preferential Cu2+ coordination by His96 and His111 induces beta-sheet formation in the unstructured amyloidogenic region of the prion protein", by C. E. Jones, S. R. Abdelraheim, D. R. Brown, J. H. Viles, Journal of Biological Chemistry, July 30 2004, 279, 31, 32018 - 32027
"From Microbes to Prions: The Final Proof of the Prion Hypothesis", by W-Q. Zou, P. Gambetti, Cell, April 22 2005, 121, 2, 155 - 162
Just as the postulates by Koch are used to determine if a specific bacterium cause a disease, so criteria must be used to determine if Prions cause a disease. The authors discuss the "prion versio of Koch's pustulates": the original disease must be reproduced in a recipient from prions grown and purified in vitro after being obtained from a donor. The PMCA ( Protein Misfolding cyclic Amplificaton) sonification cycles a fragment the putatively disease causing PrPSc and serve as new nucleation sites to convert (normal) PrPC into disease forming PrPSc. This can amplify newly formed PrPSc, reminiscent of PCR.
a "In Vitro generation of infectious scrapie Prions", by J. Castilla, P. Saa, C. Hetz, C. Soto, Cell, April 22 2005, 121, 2, 195 - 206
"Mechanisms of amyloid fibril self-assembly and inhibition Model short peptides as a key research tool", by E. Gazit, Federation of European Biochemical Societes (FEBS) Journal, Dec. 2005, 272, 23, 5971 - 5978
Amyloid fibrils are associated with twenty known syndromes, including Alzheimer's disease, Parkinson's disease, Huntington's disease, prion disorders, type II diabetes. Amyloid fibrils also perform pysiological roles with micro-organisms such as the formation of biofilms and aerial hyphae a, b, c. Thus the use of short peptide fragments that can induce the amyloid fibril state would greatly aid the study of amyloid fibril molecular recognition and self-assembly. This paper is a study of the use of short peptides that may be used to study of amyloid fibrils, as well as the effects of aromatics upon amyloid fibrils. The author notes that nano-ordered assemblies of amyloid fibrils may simply represent a state of efficient minimal energy arrangement of polypeptide chains as observed in crystalline organic and inorganic materials. In particular, the author quotes other researchers stating that amyloid fibrils are "one-dimensional crystals". Thus the author collects supporting information that short polypeptide chains have been found to induce amyloid fibril formation as a very useful experimental tool for molecular recognition and self-assembly.
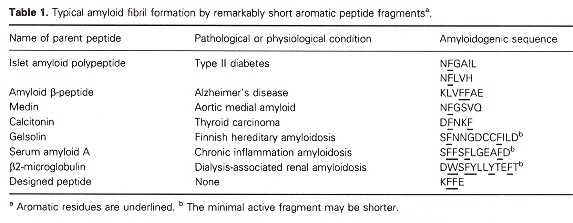

The author also points out that Short polypeptide chains that induce amyloid fibril formation have a high occurrence of aromatic residues (see the table above). This is especially provoking as aromatics are among the less frequently encountered amino acids found in proteins. The author points out that aromatic (as well as non-aromatic) peptides are found in prion formation. However, the non-aromatic peptides are much larger and take much longer timescales to form. Thus the suggestion that aromatics (such as those below) can significantly aid amyloid fibril formation in experimental studies.
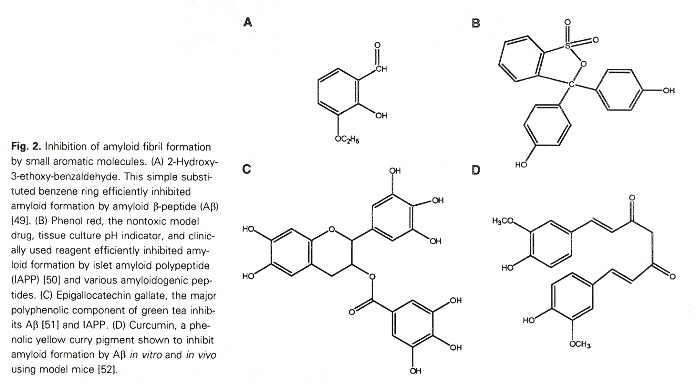
a "Role of Escherichia coli curli operons in directing amyloid fiber formation", by M. R. Chapman, L. S. Robinson, J. S. Pinker, R. Roth, J. Hauser, M. Hammar, S. Normark, S. J. Hultgren, Science, 2005, 295, 851 - 855
b "A novel class of secreted hydrophobic proteins is involved in aerial hyphae formation in Streptomyces coelicolor by forming amyloid-like fibrils", by D. Claessen, R. Rink, W. de Jong, J. Siebring, P, de Vreugd, F. G. Boersma, L. Dijkhuizen, H. A. Wosten, Genes Development, 2003, 17, 1714 - 1726
c "Amyloids - A functional coat for microorganisms", by M. F. Gebbink, D. Claessen, B. Bouma, L. Dijkhuizen, H. A. Wosten, National Review of Microbiology, 2005, 3, 333 - 341